
Ron Bose, MD, PhD
Associate Professor of Medicine
- Phone: 314-747-1171
- Fax: 314-362-7086
- Email: rbose@nospam.wustl.edu
Address:
Division of Oncology
Mail Stop 8076-0041-03
Washington University
660 South Euclid Avenue
St. Louis, MO 63110
Admin:
Rabeeka Rajput
rabeeka@wustl.edu
- Her2-positive breast cancer
- Breast cancer
- Her2/neu tyrosine kinase
- Genomics of breast cancer
- Use of proteomics to study breast cancer cell signaling
Breast cancer is typically divided into 3 major subtypes (hormone receptor positive, HER2 positive, and triple negative breast cancer). However, new genomics research has provided a new classification system for breast cancer (Luminal A, Luminal B, Basal-like, and HER2-enriched breast cancer). This research is showing that there are different spectrum of mutations and genetic alterations for each of these new categories. Basal-like breast cancer, a category that overlaps substantially with triple negative breast cancer, is characterized by an extremely high rate of mutations in a critical tumor suppressor gene, p53. This characteristic is shared by ovarian cancer and comparison of basal-like breast cancer and ovarian cancer suggests they share a lot of similarities. Hormonal receptor positive breast cancers can be divided into the new categories, Luminal A and Luminal B, with the Luminal A patients having a better prognosis. The mutation patterns in Luminal A versus B patients provide some understanding as to why this is so. HER2-enriched breast cancers are closely related to the prior category of HER2 positive breast cancers, but there are still some questions in the breast cancer field as to how a few patients can be “HER2-enriched” without having HER2 gene amplification (the defining characteristic of HER2 positive breast cancers). This is an active area of research. These findings have been published in the scientific journal, Nature, and have been described in the New York Times and the St. Louis Post-Dispatch (front pages articles in both papers on Sept 24, 2012). Dr. Bose and his lab have played an important part in developing this genomic understand of breast cancer.
A major goal of breast cancer genomics is to identify new gene targets in breast cancer for drug treatment and over 40 new possible gene targets have been identified by breast cancer genomics. Dr. Bose and his lab are taking these results forward to produce new breast cancer treatments for patients. Broadly, his research takes a 3 step approach: 1) Identify promising gene targets based on computer analysis of breast cancer genomes. 2) Perform direct testing of the genes and the proteins that they produce in the laboratory. 3) Launch clinical trials for the most promising gene targets. This process is ongoing and Dr. Bose and 2 other Oncology faculty, Drs. Cynthia Ma and Matthew Ellis, have gotten FDA-approval to start the very first genome based clinical trial.
Dr. Bose’s lab has a major interest in breast cancer cell signaling and in particular, the Her2 tyrosine kinase, which is a major protein in these cell signaling pathways. Cancer cell signaling promotes cancer cell growth and spread and is therefore a major target in the treatment of cancers. Dr. Bose has been working on the Her2 tyrosine kinase since 2003, when he was medical oncology fellow at Johns Hopkins School of Medicine. Dr. Bose has published over 8 scientific and medical papers on Her2 and its closely related proteins. Dr. Bose’s interest in cancer cell signaling started back in 1993, when he was an MD-PhD student doing research at Memorial Sloan-Kettering Cancer Center (MSKCC) in New York City. MSKCC has an affiliation with Cornell University and the joint Cornell University-MSKCC-Rockefeller University MD-PhD program, which Dr. Bose is a graduate of, is one of the top 10 MD-PhD programs in the nation.
Dr. Bose’s current research project include: 1) Studying breast cancer genomic alterations that affect cancer cell signaling. 2) Studying the biochemistry of the Her2 tyrosine kinase to determine if there are new ways to target it with drugs. 3) Study the development of resistance to current Her2 targeted drugs, such as the monoclonal antibody Herceptin, because the development of drug resistance in patient’s cancers creates a very large medical problem for patient treatment.
Dr. Bose sees patients one day each week at the Siteman Cancer Center at Barnes-Jewish West County Hospital. He sees patients with all types of breast cancer. He is particularly interested in patients with HER2-positive breast cancer and provides them information on the latest clinical trials for HER2.
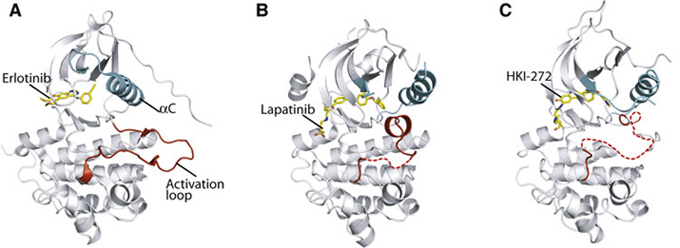
(A) Erlotinib–EGFR structure; (B) Lapatinib–EGFR structure; (C) HKI-272–EGFR structure. The panels show the entire EGFR kinase domain with inhibitor bound, revealing the active (A) and inactive (B and C) conformations of EGFR. From: Bose R, Zhang X. The ErbB kinase domain: structural perspectives into kinase activation and inhibition. Exp Cell Res 2009 Feb 15;315(4):649-58
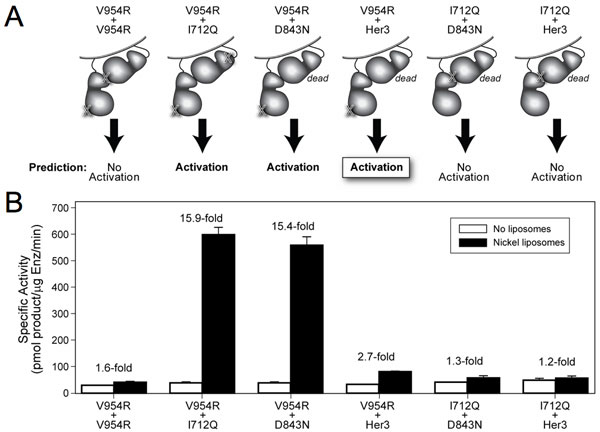
(A) Predicted effects of the Her4 mutations and of Her3 on Her4 kinase activity. The mutations shown here are in the Her4 kinase domain. (B) Her4 and Her3 kinase domain proteins were mixed at 1:1 molar ratio, liposomes containing nickel-chelating lipids were added and kinase assay performed based on phosphotransfer to an exogenous peptide substrate. From: Monsey J, Shen W, Schlesinger P, Bose R. Her4 and Her2/neu tyrosine kinase domains dimerize and activate in a reconstituted in vitro system. J Biol Chem 2010 Mar 5;285(10):7035-44
Biosketch
Education
- 1999: MD, Cornell University, Weill Medical College, New York, NY
- 1998: PhD in Pharmacology and Biochemistry, Cornell University, Graduate School of Medical Sciences, New York, NY
- Research performed at Memorial Sloan Kettering Cancer Center, New York, NY
- 1991: BS in Zoology, BA in Chemistry (with Highest Distinction), University of Rhode Island, Kingston, RI
Post-Graduate Training
- 2007-2003: Postdoctoral Research, Department of Pharmacology (Mentor: Philip Cole), Johns Hopkins University School of Medicine, Baltimore, MD
- 2007-2002: Fellowship in Medical Oncology, Department of Oncology, The Johns Hopkins Hospital, Baltimore, MD
- 2002-1999: Internship and Residency in Internal Medicine, Washington University School of Medicine in St. Louis and Barnes-Jewish Hospital, St. Louis, MO
Academic Positions
- present-2016: Associate Professor of Medicine, Departments of Medicine and Cell Biology & Physiology, Washington University School of Medicine, St. Louis, MO
- 2016-2007: Assistant Professor of Medicine, Departments of Medicine and Cell Biology & Physiology, Washington University School of Medicine, St. Louis, MO
Professional Appointments & Committees
- 2022-2020: American Association for Cancer Research (AACR) NextGen Grants for Transformative Cancer Research Scientific Review Committee
- 2019: Ad hoc Reviewer, Department of Defense, CDMRP, Breast Cancer Research Program, Clinical and Experimental Therapeutics-4 Review Panel
- 2019-2017: Medical Scientist Training Program (MSTP) Admission Committee, Washington University
- 2018: Reviewer for NIH-NCI Women Malignancy Branch
- 2018: San Antonio Breast Cancer Symposium (SABCS), Abstract Reviewer for the SABCS Program Committee
- 2018-2015: Standing Member (permanent member), American Cancer Society, Tumor Biology and Genomics Committee
- 2017: Ad hoc Reviewer for NCI Subcommittee on Transition to Independence Grants (K99/R00 and K22)
- 2014: Reviewer for NIH-NCI Center for Cancer Research Site Visit
- 2014-2012: Grant Reviewer, American Cancer Society, Tumor Biology and Genomics Committee
- 2013-2012: Member of the AACR Education Committee for the 2013 AACR Annual Meeting
- 2013-2011: Member of Writing Committee, NCI – The Cancer Genome Atlas (TCGA) Lung Cancer Project
Editorial Boards
- present-2015: Editorial Board Member, Breast Cancer Research
Board Certification
- 2014, 2004: Diplomate, American Board of Internal Medicine, Medical Oncology (active)
- 2002: Diplomate, American Board of Internal Medicine, Internal Medicine (lapsed)
Honors & Awards
- 2017: Elected to the American Society for Clinical Investigation (ASCI)
- 2014-2013: Susan G. Komen for the Cure, St. Louis Affiliate, “Pink Tie Guy”
- 2013: Susan G. Komen for the Cure, St. Louis, Health Professional of the Year
- 2013: Best Paper of the Year in the Journal of Biological Chemistry in the Category of Signal Transduction
- 2006: AACR Scholar-in-Training Travel Award
- 2003: Selected as Senior Resident to give Grand Rounds to Department of Medicine, Washington University School of Medicine
- 2002: Resident’s Teaching Award, Washington University
- 2000, 1999: Caring Spirit Award, Barnes-Jewish Hospital
- 1997, 1995: Award of Excellence, Vincent du Vigneaud Symposium, Cornell University
- 1993: Alpha Omega Alpha Medical Honor Society
Professional Memberships
- American Society for Clinical Investigation
- American Society of Clinical Oncology
- American Association for Cancer Research