
Matthew J. Walter, MD
Edward P. Evans Endowed Professor of Myelodysplastic Syndromes; Section Director – Stem Cell Biology
- Phone: 314-454-8304
- Fax: 314-454-7551
- Email: mjwalter@nospam.wustl.edu
Address:
Oncology Division
Mail Stop 8007-0057-06
Washington University
660 South Euclid Avenue
St. Louis, MO 63110
Admin:
Melinda Dioneda
dionedal@wustl.edu
- Myelodysplastic syndromes
- Acute myeloid leukemia
- Functional genomics
- Tumor biology
- Hematopoiesis
The main interest of our laboratory is the discovery and characterization of gene mutations in patients with myelodysplastic syndrome (MDS) with the long-term goal of understanding the molecular mechanisms that control abnormal hematopoiesis. MDS results from the expansion of one or more dominant hematopoietic clones that contain initiating somatic mutations, while transformation from MDS to acute myeloid leukemia (AML) occurs as these clones accumulate additional progression factors (including point mutations in genes and cytogenetic abnormalities).
Research is ongoing in the following areas:
- MDS genomics and tumor clonal heterogeneity
Gene mutations are identified in the genomes of hematopoietic cells from patients with MDS and AML using next generation sequencing technology, including whole genome sequencing. Somatic mutations are used to decipher the clonal architecture of MDS and AML samples (ie, founding clones and subclones). The functional significance of recurrently mutated genes is then studied using primary human hematopoietic cells and mouse models to understand the mechanisms of disease pathogenesis. - The role of splicing gene mutations in MDS pathogenesis
Ongoing projects are focused on understanding the mechanism of MDS initiation and progression induced by mutations in genes that participate in pre-mRNA splicing identified by our group and others. Spliceosome gene mutations occur in 50% of MDS patients. Functional splicing assays, next-generation sequencing technology, and hematopoietic assays will be used to study primary human cells and mouse models harboring mutations. Preclinical cell and mouse models are being used to test novel approaches to selectively kill spliceosome mutant cells with the goal of introducing new therapies into clinical trials for MDS patients. - Monitoring tumor burden using next-generation sequencing
Gene mutations are identified in patient samples using whole genome and exome sequencing. The abundance of tumor-specific mutations are tracked in serial samples using error-corrected sequencing during treatment to objectively monitor tumor burden and treatment response. The prognostic utility of these approaches are being tested in clinical trials to improve patient outcomes.
All these studies involve state of the art technologies and incorporate functional genomics, bioinformatics, next-generation sequencing, stem cell and tumor biology, and mouse models of disease to study hematopoiesis.
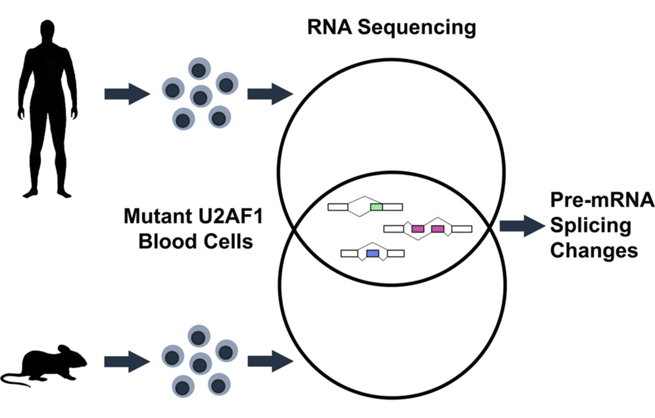
Shirai et al. show that the U2AF1 mutation most commonly found in MDS alters pre-mRNA splicing in RNA processing genes, ribosomal genes, and recurrently mutated MDS and acute myeloid leukemia-associated genes in hematopoietic progenitor cells and affect hematopoiesis.
From: Mutant U2AF1 expression alters hematopoiesis and pre-mRNA splicing in vivo.
Cancer Cell 2015 May 11;27(5):631-43
Biosketch
Education
- 1995: MD, St. Louis University School of Medicine, St. Louis, MO
- 1990: BS, American University, Washington, DC
Post-Graduate Training
- 2004-2001: Post-Doctoral Fellow, Washington University, St. Louis, MO (Timothy Ley, MD, Mentor)
- 2004-2000: Hematology-Oncology Fellow, Washington University Medical Center, St. Louis, MO
- 2000-1999: Assistant Chief of Service (Chief Medical Resident), Johns Hopkins Hospital, Baltimore, MD
- 1999-1998: Hematology-Oncology Fellow, Washington University Medical Center, St. Louis, MO
- 1998-1996: Resident in Medicine, Johns Hopkins Hospital, Baltimore, MD
- 1996-1995: Intern in Medicine, Johns Hopkins Hospital, Baltimore, MD
- 1990: BS, American University, Washington, DC
Academic Positions
- present-2020: Edward P. Evans Endowed Professor of Myelodysplastic Syndromes
- present-2019: Director of the Edward P. Evans Center for MDS, Washington University School of Medicine, St. Louis, MO
- present-2017: Professor of Medicine and Genetics, Washington University School of Medicine, St. Louis, MO
- 2017-2019: Scientific Director of the MDS Program, Washington University School of Medicine, St. Louis, MO
- 2013-2017: Associate Professor of Medicine and Genetics, Washington University School of Medicine, St. Louis, MO
- 2013-2005: Assistant Professor of Medicine and Genetics, Washington University School of Medicine, St. Louis, MO
- 2005-2004: Instructor in Medicine, Washington University School of Medicine, St. Louis, MO
University & Hospital Appointments & Committees
- present-2019: Member, Molecular Genetics and Genomics DBBS Steering Committee at Washington University
- 2019: Member, Search Committee for Chief of Hematology at Washington University
- 2018: Member, Search Committee for Chief of Pediatric Hematology Oncology at Washington University
- 2013-2007: Co-course Master, Lucille P. Markey Special Emphasis Pathway in Human Pathobiology, Division of Biology and Biomedical Sciences
Board Certification
- 2002: American Board of Internal Medicine, Hematology (active)
- 2012-2002: American Board of Internal Medicine, Oncology (inactive)
- 2008-1998: American Board of Internal Medicine, Internal Medicine (inactive)
Honors & Awards
- 2020: Elected, Association of American Physicians
- 2018-2013: Leukemia and Lymphoma Society Scholar Award
- 2013: Elected, American Society for Clinical Investigation
- 2009-2003: NHLBI Loan Repayment Program recipient
- 2007-2005: American Society of Hematology Scholar Award
- 1995: Medical Degree with Distinction in Research, St. Louis University School of Medicine
- 1994: Alpha Omega Alpha Honor Medical Society, St. Louis University School of Medicine
- 1993-1992: Howard Hughes Medical Institute – National Institutes of Health Research Scholar, Bethesda, MD
Editorial Responsibilities
- 2015-2012: Editorial Board Member, Leukemia
National Scientific Panels
- present-2015: American Society of Hematology (ASH) Task Force on Precision Medicine
- present-2014: Member, The National MDS Study Protocol Team, NIH
- 2017-2014: American Society of Hematology (ASH) Scientific Committee on Myeloid Neoplasia
- 2013: American Society of Hematology (ASH) Scientific Committee on Myeloid Biology
- 2013: American Association for Cancer Research (AACR) Annual Meeting Education Committee
Professional Societies
- American Society of Hematology
- America Society for Clinical Investigation
- Research Member, Siteman Cancer Center, Washington University
- Association of American Physicians